Sulfur Starvation
| Portal |
|---|
|
Yeast Sulfur Project Portal |
| Home |
|
Sulfur Starvation Homepage |
| Supplement |
|
supplementary figures and tables |
| Download |
|
raw and derived data used in the publication |
| Figures |
|
from the paper |
| Authors |
|
contacts who worked on the project |
| Supplement | |
|---|---|
|
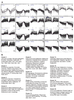
Figure S1. k-means clustering of Met-dependent genes in two biosynthetic mutants.
A) k-means clustering and B) Gene Ontology (GO) term enrichment for genes that are similarly expressed in met6Δ and met13Δ during Met starvation. Each cluster shows data for met6Δ (left) and met13Δ (right). |

|
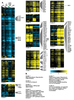
Figure S2. All genes whose expression depends on MET31/MET32.
A) All genes that depend statistically significantly on MET31/MET32 but not CBF1. Strain order is met6Δ, cbf1Δmet6Δ, met31Δmet32Δmet6Δ, met4Δ. Genes of particular interest are shown. B) GO term enrichment. |

|
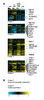
Figure S3. All genes whose expression depends on CBF1.
A) All genes that depend statistically significantly on CBF1 but not MET31/MET32. Strain order is met6Δ, cbf1Δmet6Δ, met31Δmet32Δmet6Δ, met4Δ. Genes of particular interest are shown. B) GO term enrichment. |

|
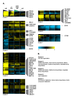
Figure S4. All genes whose expression depends on MET31/MET32 and CBF1.
A) All genes that depend statistically significantly on MET31/MET32 and CBF1. Strain order is met6Δ, cbf1Δmet6Δ, met31Δmet32Δmet6Δ, met4Δ. Genes of particular interest are shown. B) GO term enrichment. |

|
Figure S5. Activity heatmap for all transcription factors.
Hierarchically-clustered heat-map representation of the calculated activities for all TFs. |
|
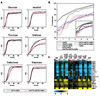
Figure S6.
A) Comparison of cbf1Δmet6Δ (black) and met31Δmet32Δmet6Δ (red) growth rates in different carbon sources. B) Cell cycle arrest phenotypes. Percent unbudded cells (i.e. bud index) measured during six hours of Met starvation for three replicates each of met6Δ, which arrests efficiently, and cbf1Δmet31Δmet6Δ and cbf1Δmet32Δmet6Δ, which do not arrest efficiently. Inset: Bud index for all strains. C) Potentially Met-dependent cell cycle genes. |
|
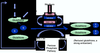
Figure S7.
Relationship between antioxidants and Met-regulated metabolites. |